In most of the real-life cases when we would be representing or modeling some event, we would be dealing with a lot of random variables. Even if we would consider all the random variables to be discrete, there would still be exponentially large number of values in the joint probability distribution. Dealing with such huge amount of data would be computationally expensive (and in some cases, even intractable), and would also require huge amount of memory to store the probability of each combination of states of these random variables.
However, in most of the cases, many of these variables are marginally or conditionally independent of each other. By exploiting these independencies, we can reduce the number of values we need to store to represent the joint probability distribution.
For instance, in the previous restaurant example, the joint probability distribution across the four random variables that we discussed (that is, quality of food Q, location of restaurant L, cost of food C, and the number of people visiting N) would require us to store 23 independent values. By the chain rule of probability, we know the following:
P(Q, L, C, N) = P(Q) P(L|Q) P(C|L, Q) P(N|C, Q, L)
Now, let us try to exploit the marginal and conditional independence between the variables, to make the representation more compact. Let's start by considering the independency between the location of the restaurant and quality of food over there. As both of these attributes are independent of each other, P(L|Q) would be the same as P(L). Therefore, we need to store only one parameter to represent it. From the conditional independence that we have seen earlier, we know that ![]() . Thus, P(N|C, Q, L) would be the same as P(N|C, L); thus needing only four parameters. Therefore, we now need only (2 + 1 + 6 + 4 = 13) parameters to represent the whole distribution.
. Thus, P(N|C, Q, L) would be the same as P(N|C, L); thus needing only four parameters. Therefore, we now need only (2 + 1 + 6 + 4 = 13) parameters to represent the whole distribution.
We can conclude that exploiting independencies helps in the compact representation of joint probability distribution. This forms the basis for the Bayesian network.
A Bayesian network is represented by a Directed Acyclic Graph (DAG) and a set of Conditional Probability Distributions (CPD) in which:
In our previous restaurant example, the nodes would be as follows:
- Quality of food (Q)
- Location (L)
- Cost of food (C)
- Number of people (N)
As the cost of food was dependent on the quality of food (Q) and the location of the restaurant (L), there will be an edge each from Q → C and L → C. Similarly, as the number of people visiting the restaurant depends on the price of food and its location, there would be an edge each from L → N and C → N. The resulting structure of our Bayesian network is shown in Fig 1.3:
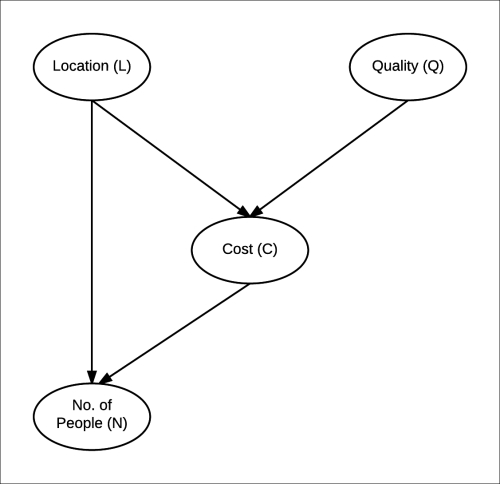
Fig 1.3: Bayesian network for the restaurant example
Each node in our Bayesian network for restaurants has a CPD associated to it. For example, the CPD for the cost of food in the restaurant is P(C|Q, L), as it only depends on the quality of food and location. For the number of people, it would be P(N|C, L) . So, we can generalize that the CPD associated with each node would be P(node|Par(node)) where Par(node) denotes the parents of the node in the graph. Assuming some probability values, we will finally get a network as shown in Fig 1.4:
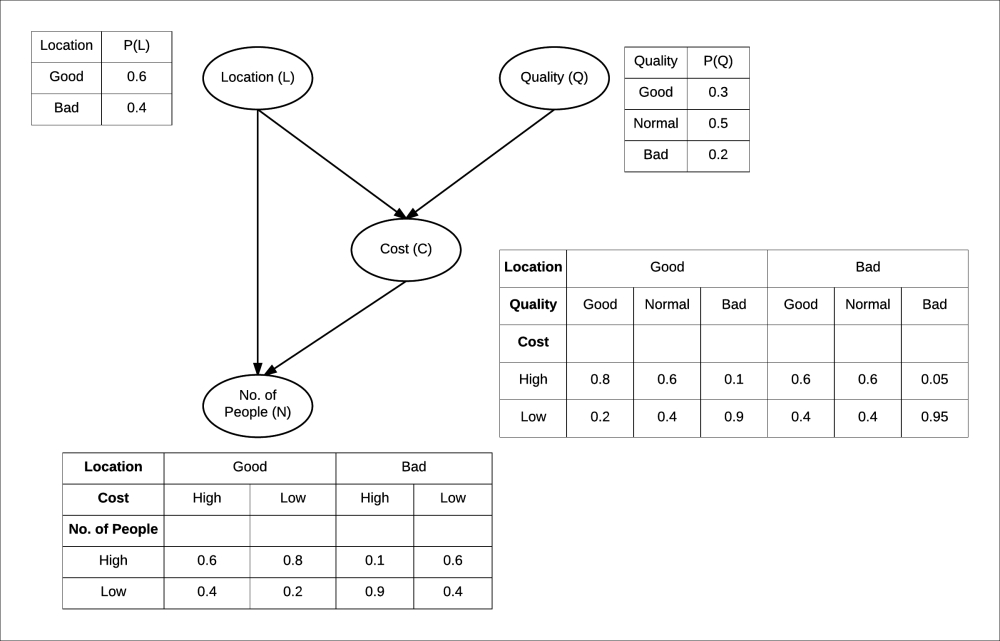
Fig 1.4: Bayesian network of restaurant along with CPDs
Let us go back to the joint probability distribution of all these attributes of the restaurant again. Considering the independencies among variables, we concluded as follows:
P(Q,C,L,N) = P(Q)P(L)P(C|Q, L)P(N|C, L)
So now, looking into the Bayesian network (BN) for the restaurant, we can say that for any Bayesian network, the joint probability distribution ![]() over all its random variables
over all its random variables ![]() can be represented as follows:
can be represented as follows:
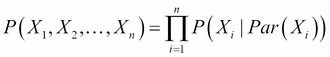
This is known as the chain rule for Bayesian networks.
Also, we say that a distribution P factorizes over a graph G, if P can be encoded as follows:
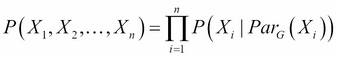
Here, ![]() is the parent of X in the graph G.
is the parent of X in the graph G.
Let us consider a more complex Bayesian network of a student getting late for school, as shown in Fig 1.5:
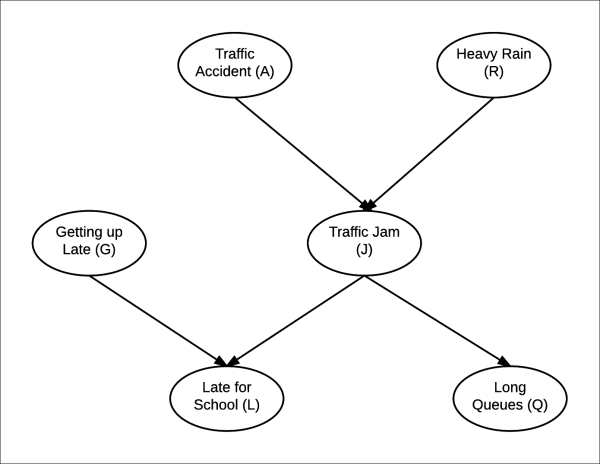
Fig 1.5: Bayesian network representing a particular day of a student going to school
For this Bayesian network, just for simplicity, let us assume that each random variable is discrete with only two possible states {yes, no}.
In pgmpy, we can initialize an empty BN or a model with nodes and edges. We can initializing an empty model as follows:
In [1]: from pgmpy.models import BayesianModel In [2]: model = BayesianModel()
We can now add nodes and edges to this network:
In [3]: model.add_nodes_from(['rain', 'traffic_jam'])
In [4]: model.add_edge('rain', 'traffic_jam')If we add an edge, but the nodes, between which the edge is, are not present in the model, pgmpy automatically adds those nodes to the model.
In [5]: model.add_edge('accident', 'traffic_jam')
In [6]: model.nodes()
Out[6]: ['accident', 'rain', 'traffic_jam']
In [7]: model.edges()
Out[7]: [('rain', 'traffic_jam'), ('accident', 'traffic_jam')]In the case of a Bayesian network, each of the nodes has an associated CPD with it. So, let's define some tabular CPDs to associate with the model:
In [8]: from pgmpy.factors import TabularCPD
In [9]: cpd_rain = TabularCPD('rain', 2, [[0.4], [0.6]])
In [10]: cpd_accident = TabularCPD('accident', 2, [[0.2], [0.8]])
In [11]: cpd_traffic_jam = TabularCPD(
'traffic_jam', 2,
[[0.9, 0.6, 0.7, 0.1],
[0.1, 0.4, 0.3, 0.9]],
evidence=['rain', 'accident'],
evidence_card=[2, 2])Here, we defined three CPDs. We now need to associate them with our model. To associate them with the model, we just need to use the add_cpd method and pgmpy automatically figures out which CPD is for which node:
In [12]: model.add_cpds(cpd_rain, cpd_accident, cpd_traffic_jam)
In [13]: model.get_cpds()
Out[13]:
[<TabularCPD representing P(rain:2) at 0x7f477b6f9940>,
<TabularCPD representing P(accident:2) at 0x7f477b6f97f0>,
<TabularCPD representing P(traffic_jam:2 | rain:2, accident:2) at
0x7f477b6f9e48>]Now, let's add the remaining variables and their CPDs:
In [14]: model.add_node('long_queues')
In [15]: model.add_edge('traffic_jam', 'long_queues')
In [16]: cpd_long_queues = TabularCPD('long_queues', 2,
[[0.9, 0.2],
[0.1, 0.8]],
evidence=['traffic_jam'],
evidence_card=[2])
In [17]: model.add_cpds(cpd_long_queues)
In [18]: model.add_nodes_from(['getting_up_late',
'late_for_school'])
In [19]: model.add_edges_from(
[('getting_up_late', 'late_for_school'),
('traffic_jam', 'late_for_school')])
In [20]: cpd_getting_up_late = TabularCPD('getting_up_late', 2,
[[0.6], [0.4]])
In [21]: cpd_late_for_school = TabularCPD(
'late_for_school', 2,
[[0.9, 0.45, 0.8, 0.1],
[0.1, 0.55, 0.2, 0.9]],
evidence=['getting_up_late',
'traffic_jam'],
evidence_card=[2, 2])
In [22]: model.add_cpds(cpd_getting_up_late, cpd_late_for_school)
In [23]: model.get_cpds()
Out[23]:
[<TabularCPD representing P(rain:2) at 0x7f477b6f9940>,
<TabularCPD representing P(accident:2) at 0x7f477b6f97f0>,
<TabularCPD representing P(traffic_jam:2 | rain:2, accident:2) at
0x7f477b6f9e48>,
<TabularCPD representing P(long_queues:2 | traffic_jam:2) at
0x7f477b7051d0>,
<TabularCPD representing P(getting_up_late:2) at 0x7f477b7059e8>,
<TabularCPD representing P(late_for_school:2 | getting_up_late:2,
traffic_jam:2) at 0x7f477b705dd8>]Additionally, pgmpy also provides a check_model method that checks whether the model and all the associated CPDs are consistent:
In [24]: model.check_model() Out[25]: True
In case we have got some wrong CPD associated with the model and we want to remove it, we can use the remove_cpd method. Let's say we want to remove the CPD associated with variable late_for_school, we could simply do as follows:
In [26]: model.remove_cpds('late_for_school')
In [27]: model.get_cpds()
Out[27]:
[<TabularCPD representing P(rain:2) at 0x7f477b6f9940>,
<TabularCPD representing P(accident:2) at 0x7f477b6f97f0>,
<TabularCPD representing P(traffic_jam:2 | rain:2, accident:2) at
0x7f477b6f9e48>,
<TabularCPD representing P(long_queues:2 | traffic_jam:2) at
0x7f477b7051d0>,
<TabularCPD representing P(getting_up_late:2) at 0x7f477b7059e8>]Would the probability of having a road accident change if I knew that there was a traffic jam? Or, what are the chances that it rained heavily today if some student comes late to class? Bayesian networks helps in finding answers to all these questions. Reasoning patterns are key elements of Bayesian networks.
Before answering all these questions, we need to compute the joint probability distribution. For ease in naming the nodes, let's denote them as follows:
- Traffic accident as A
- Heavy rain as B
- Traffic jam as J
- Getting up late as G
- Long queues as Q
- Late to school as L
From the chain rule of the Bayesian network, we have the joint probability distribution ![]() as follows:
as follows:
Starting with a simple query, what are the chances of having a traffic jam if I know that there was a road accident? This question can be put formally as what is the value of P(J|A = True)?
First, let's compute the probability of having a traffic jam P(J). P(J) can be computed by summing all the cases in the joint probability distribution, where J = True and J = False, and then renormalize the distribution to sum it to 1. We get P(J = True) = 0.416 and P(J = True) = 0.584.
To compute P(J|A = True), we have to eliminate all the cases where A = False, and then we can follow the earlier procedure to get P(J|A = True). This results in P(J = True|A = True) = 0.72 and P(J = False|A = True) = 0.28. We can see that the chances of having a traffic jam increased when we knew that there was an accident. These results match with our intuition. From this, we conclude that the observation of the outcome of the parent in a Bayesian network influences the probability of its children. This is known as causal reasoning. Causal reasoning need not only be the effect of parent on its children; it can go further downstream in the network.
We have seen that the observation of the outcome of parents influence the probability of the children. Is the inverse possible? Let's try to find the probability of heavy rain if we know that there is a traffic accident. To do so, we have to eliminate all the cases where J = False and then reduce the probability to get P(R|J = True). This results in P(R = True|J = True) = 0.7115 and P(R = False|J = True) = 0.2885. This is also intuitive. If we knew that there was a traffic jam, then the chances of heavy rain would increase. This is known as evidential reasoning, where the observation of the outcomes of the children or effect influences the probability of parents or causes.
Let's look at another type of reasoning pattern. If we knew that there was a traffic jam on a day when there was no heavy rain, would it affect the chances of a traffic accident? To do so, we have to follow a similar procedure of eliminating all those cases, except the ones where R = False and J = True. By doing so, we would get P(A = True|J = True, R = False) = 0.6 and P(A = False|J = True, R = False) = 0.4. Now, the probability of an accident increases, which is what we had expected. As we can see that before the observation of the traffic jam, both the random variables, heavy rain and traffic accident, were independent of each other, but with the observation of their common children, they are now dependent on each other. This type of reasoning is called as intercausal reasoning, where different causes with the same effect influence each other.
In the last section, we saw how influence flows in a Bayesian network, and how observing some event changes our belief about other variables in the network. In this section, we will discuss the independence conditions that hold in a Bayesian network no matter which probability distribution is parameterizing the network.
In any network, there can be two types of connections between variables, direct or indirect. Let's start by discussing the direct connection between variables.
In the case of direct connections, we have a direct connection between two variables, that is, there's an edge X → Y in the graph G. In the case of a direct connection, we can always find some probability distribution where they are dependent. Hence, there is no independence condition in a direct connection, no matter which other variables are observed in the network.
In the case of indirect connections, we have four different ways in which the variables can be connected. They are as follows:
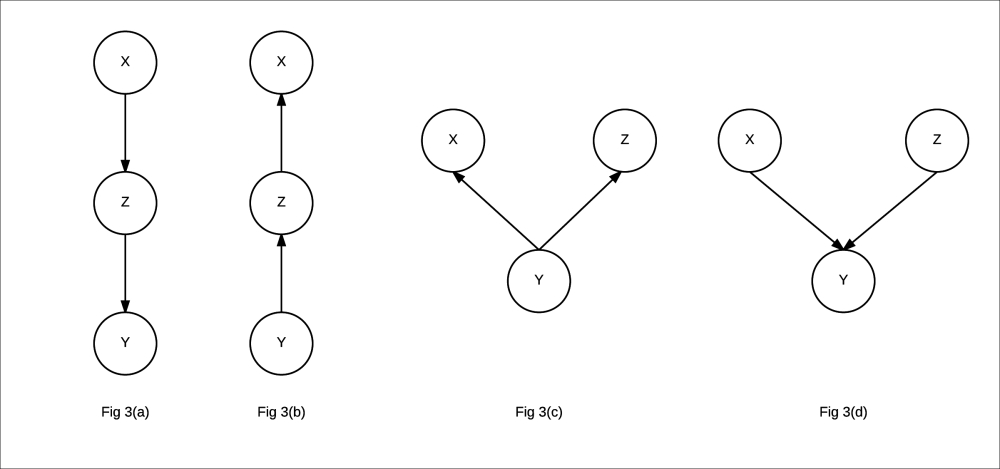
Fig 3(a): Indirect causal relationship
Fig 3(b): Indirect evidential relationship
Fig 3(c): Common cause relationship
Fig 3(d): Common effect relationship
- Indirect causal effect: Fig 3(a) shows an indirect causal relationship between variables X and Y. For intuition, let's consider the late-for-school model, where A → J →L is a causal relationship. Let's first consider the case where J is not observed. If we observe that there has been an accident, then it increases our belief that there would be a traffic jam, which eventually leads to an increase in the probability of getting late for school. Here we see that if the variable J is not observed, then A is able to influence L through J. However, if we consider the case where J is observed, say we have observed that there is a traffic jam, then irrespective of whether there has been an accident or not, it won't change our belief of getting late for school. Therefore, in this case we see that
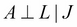 .
More often, in the case of an indirect causal relationship
.
More often, in the case of an indirect causal relationship 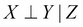 .
.
- Indirect evidential effect: Fig 3(b) represents an indirect evidential relationship. In the late-for-school model, we can again take the example of L → J ← A. Let's first take the case where we haven't observed J. Now, if we observe that somebody is late for school, it increases our belief that there might be a traffic jam, which increases our belief about there being an accident. This leads us to the same results as we got in the case of an indirect causal effect. The variables X and Y are dependent, but become independent if we observe Z, that is
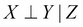 .
.
- Common cause: Fig 3(c) represents a common cause relationship. Let's take the example of L ← J → Q from our late-for-school model. Taking the case where J is not observed, we see that getting late for school makes our belief of being in a traffic jam stronger, which also leads to an increase in the probability of being in a long queue. However, what if we already have observed that there was a traffic jam? In this case, getting late for school doesn't have any effect on being in a long queue. Hence, we see that the independence conditions in this case are also the same as we saw in the previous two cases, that is, X is able to influence Y through Z only if Z is not observed.
- Common effect: Fig 3(d) represents a common effect relationship. Taking the example of A → J ← B from the late-for-school model, if we have an observation that there was an accident, it increases the probability of having a traffic jam, but does not have any effect on the probability of heavy rain. Hence, A | B. We see that we have a different observation here than the previous three cases. Now, if we consider the case when J is observed, let's say that there has been a jam. If we now observe that there hasn't been an accident, it does increase the probability that there might have been heavy rain. Hence, A is not independent of B if J is observed. More generally, we can say that in the case of common effect, X is independent of Y if, and only if, Z is not observed.
Now, in a network, how do we know if a variable influences another variable? Let's say we want to check the independence conditions for ![]() and
and ![]() . Also, let's say they are connected by a trail
. Also, let's say they are connected by a trail ![]() and let Z be the set of observed variables in the Bayesian network. In this case,
and let Z be the set of observed variables in the Bayesian network. In this case, ![]() will be able to influence
will be able to influence ![]() if and only if the following two conditions are satisfied:
if and only if the following two conditions are satisfied:
-
For every V structure of the form
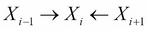 in the trail, either
in the trail, either 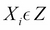 or any descendant of
or any descendant of 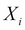 is an element of Z
is an element of Z - No other node on the trail is in Z
Also, if an influence can flow in a trail in a network, it is known as an active trail. Let's see some examples to check the active trails using pgmpy for the late-for-school model:
In [28]: model.is_active_trail('accident', 'rain')
Out[28]: False
In [29]: model.is_active_trail('accident', 'rain',
observed='traffic_jam')
Out[29]: True
In [30]: model.is_active_trail('getting_up_late', 'rain')
Out[30]: False
In [31]: model.is_active_trail('getting_up_late', 'rain',
observed='late_for_school')
Out[31]: True