The Local Outlier Factor (LOF) is an outlier detection algorithm that detects the outliers based on comparing the local density of the data instance with its neighbors. It does this in order to decide if the data instance belongs to a region of similar density. It can detect an outlier in a dataset, in such circumstances where the number of clusters are unknown and the clusters are of different density and sizes. It's inspired by the KNN (K-Nearest Neighbors) algorithm and is widely used.
In the previous recipe, we looked at univariate data. In this one, we will use multivariate data and try to find outliers. Let's use a very small dataset to understand the LOF algorithm for outlier detection.
We will create a 5 X 2 matrix, and looking at the data, we know that the last tuple is an outlier. Let's also plot it as a scatter plot:
from collections import defaultdict import numpy as np instances = np.matrix([[0,0],[0,1],[1,1],[1,0],[5,0]]) import numpy as np import matplotlib.pyplot as plt x = np.squeeze(np.asarray(instances[:,0])) y = np.squeeze(np.asarray(instances[:,1])) plt.cla() plt.figure(1) plt.scatter(x,y) plt.show()
The plot looks as follows:
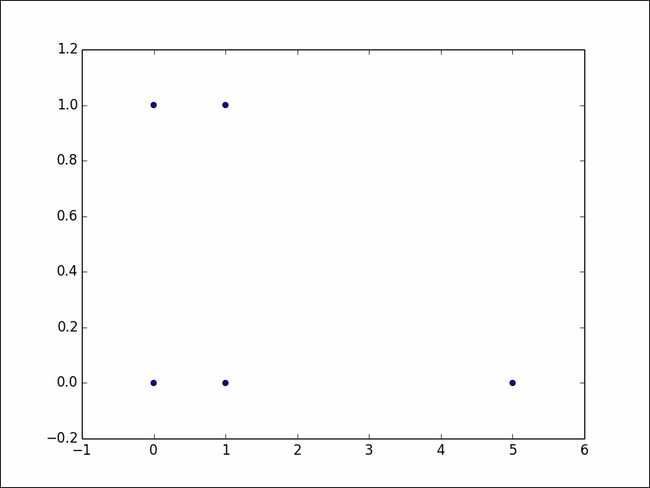
LOF works by calculating the local density of each point. Based on the distance of k-nearest neighbors of a point, the local density of the point is estimated. By comparing the local density of the point with the densities of its neighbors, outliers are detected. Outliers have a low density compared with their neighbors.
We will need to go through some term definitions in order to understand LOF:
- The k-distance of object P is the distance between the object P and its kth nearest neighbor. K is a parameter of the algorithm.
- The k-distance neighborhood of P is the list of all the objects, Q, whose distance from P is either less than or equal to the distance between P and its kth nearest object.
- The reachability distance from P to Q is defined as the maximum of the distance between P and its kth nearest neighbor, and the distance between P and Q. The following notation may help clarify this:
Reachability distance (P ß Q) = > maximum(K-Distance(P), Distance(P,Q))
- The Local Reachability Density of P (LRD(P)) is the ratio of the k-distance neighborhood of P and the sum of the reachability distance of k and its neighborhood.
- The Local Outlier Factor of P (LOF(P)) is the average of the ratio of the local reachability of P and those of P's k-nearest neighbors.
- Let's get the
pairwisedistance between the points:k = 2 distance = 'manhattan' from sklearn.metrics import pairwise_distances dist = pairwise_distances(instances,metric=distance)
- Let's calculate the k-distance. We will use
heapqand get the k-nearest neighbors:# Calculate K distance import heapq k_distance = defaultdict(tuple) # For each data point for i in range(instances.shape[0]): # Get its distance to all the other points. # Convert array into list for convienience distances = dist[i].tolist() # Get the K nearest neighbours ksmallest = heapq.nsmallest(k+1,distances)[1:][k-1] # Get their indices ksmallest_idx = distances.index(ksmallest) # For each data point store the K th nearest neighbour and its distance k_distance[i]=(ksmallest,ksmallest_idx) - Calculate the k-distance neighborhood:
def all_indices(value, inlist): out_indices = [] idx = -1 while True: try: idx = inlist.index(value, idx+1) out_indices.append(idx) except ValueError: break return out_indices # Calculate K distance neighbourhood import heapq k_distance_neig = defaultdict(list) # For each data point for i in range(instances.shape[0]): # Get the points distances to its neighbours distances = dist[i].tolist() print "k distance neighbourhood",i print distances # Get the 1 to K nearest neighbours ksmallest = heapq.nsmallest(k+1,distances)[1:] print ksmallest ksmallest_set = set(ksmallest) print ksmallest_set ksmallest_idx = [] # Get the indices of the K smallest elements for x in ksmallest_set: ksmallest_idx.append(all_indices(x,distances)) # Change a list of list to list ksmallest_idx = [item for sublist in ksmallest_idx for item in sublist] # For each data pont store the K distance neighbourhood k_distance_neig[i].extend(zip(ksmallest,ksmallest_idx)) - Then, calculate the reachability distance and LRD:
#Local reachable density local_reach_density = defaultdict(float) for i in range(instances.shape[0]): # LRDs numerator, number of K distance neighbourhood no_neighbours = len(k_distance_neig[i]) denom_sum = 0 # Reachability distance sum for neigh in k_distance_neig[i]: # maximum(K-Distance(P), Distance(P,Q)) denom_sum+=max(k_distance[neigh[1]][0],neigh[0]) local_reach_density[i] = no_neighbours/(1.0*denom_sum) - Calculate LOF:
lof_list =[] #Local Outlier Factor for i in range(instances.shape[0]): lrd_sum = 0 rdist_sum = 0 for neigh in k_distance_neig[i]: lrd_sum+=local_reach_density[neigh[1]] rdist_sum+=max(k_distance[neigh[1]][0],neigh[0]) lof_list.append((i,lrd_sum*rdist_sum))
In step 1, we select our distance metric to be Manhattan and our k value as two. We are looking at the second nearest neighbor for our data point.
We must then proceed to calculate the pairwise distance between our tuples. The pairwise similarity is stored in the dist matrix. As you can see, the shape of dist is as follows:
>>> dist.shape (5, 5) >>>
It is a 5 X 5 matrix, where the rows and columns are individual tuples and the cell value indicates the distance between them.
In step 2, we then import heapq:
import heapq
heapq is a data structure that is also known as a priority queue. It is similar to a regular queue except that each element is associated with a priority, and an element with a high priority is served before an element with a low priority.
Refer to the Wikipedia link for more information on priority queues:
http://en.wikipedia.org/wiki/Priority_queue.
The Python heapq documentation can be found at https://docs.python.org/2/library/heapq.html.
k_distance = defaultdict(tuple)
Next, we define a dictionary where the key is the tuple ID and the value is the distance of the tuple to its kth nearest neighbor. In our case, it should be the second nearest neighbor.
We then enter a for loop in order to find the kth nearest neighbor's distance for each of the data points:
distances = dist[i].tolist()
From our distance matrix, we extract the ith row. As you can see, the ith row captures the distance between the object i and all the other objects. Remember that the cell value (i,i) holds the distance to itself. We need to ignore this in the next step. We must convert the array to a list for our convenience. Let's try to understand this with an example. The distance matrix looks as follows:
>>> dist
array([[ 0., 1., 2., 1., 5.],
[ 1., 0., 1., 2., 6.],
[ 2., 1., 0., 1., 5.],
[ 1., 2., 1., 0., 4.],
[ 5., 6., 5., 4., 0.]]) Let's assume that we are in the first iteration of our for loop and hence, our i =0. (remember that the Python indexing starts with 0).
So, now our distances list will look as follows:
[ 0., 1., 2., 1., 5.]
From this, we need the kth nearest neighbor, that is, the second nearest neighbor, as we have set K = 2 at the beginning of the program.
Looking at it, we can see that both index 1 and index 3 can be our the kth nearest neighbor as both have a value of 1.
Now, we use the heapq.nsmallest function. Remember that we had mentioned that heapq is a normal queue but with a priority associated with each element. The value of the element is the priority in this case. When we say that give me the n smallest, heapq will return the smallest elements:
# Get the Kth nearest neighbours ksmallest = heapq.nsmallest(k+1,distances)[1:][k-1]
Let's look at what the heapq.nsmallest function does:
>>> help(heapq.nsmallest)
Help on function nsmallest in module heapq:
nsmallest(n, iterable, key=None)
Find the n smallest elements in a dataset.
Equivalent to: sorted(iterable, key=key)[:n]It returns the n smallest elements from the given dataset. In our case, we need the second nearest neighbor. Additionally, we need to avoid (i,i) as mentioned previously. So we must pass n = 3 to heapq.nsmallest. This ensures that it returns the three smallest elements. We then subset the list to exclude the first element (see [1:] after nsmallest function call) and finally retrieve the second nearest neighbor (see [k-1] after [1:]).
We must also get the index of the second nearest neighbor of i and store it in our dictionary:
# Get their indices ksmallest_idx = distances.index(ksmallest) # For each data point store the K th nearest neighbour and its distance k_distance[i]=(ksmallest,ksmallest_idx)
Let's print our dictionary:
print k_distance
defaultdict(<type 'tuple'>, {0: (1.0, 1), 1: (1.0, 0), 2: (1.0, 1), 3: (1.0, 0), 4: (5.0, 0)})Our tuples have two elements: the distance, and the index of the elements in the distances array. So, for instance 0, the second nearest neighbor is the element in index 1.
Having calculated the k-distance for all our data points, we then move on to find the k-distance neighborhood.
In step 3, we find the k-distance neighborhood for each of our data points:
# Calculate K distance neighbourhood import heapq k_distance_neig = defaultdict(list)
Similar to our previous step, we import the heapq module and declare a dictionary that is going to hold our k-distance neighborhood details. Let's recap what the k-distance neighborhood is:
The k-distance neighborhood of P is the list of all the objects, Q, whose distance from P is either less than or equal to the distance between P and its kth nearest object:
distances = dist[i].tolist() # Get the 1 to K nearest neighbours ksmallest = heapq.nsmallest(k+1,distances)[1:] ksmallest_set = set(ksmallest)
The first two lines should be familiar to you. We did this in our previous step. Look at the second line. Here, we invoked n smallest again with n=3 in our case (K+1), but we selected all the elements in the output list except the first one. (Guess why? The answer is in the previous step.)
Let's see it in action by printing the values. As usual, in the loop, we assume that we are seeing the first data point or tuple where i=0.
Our distances list is as follows:
[0.0, 1.0, 2.0, 1.0, 5.0]
Our
heapq.nsmallest function returns the following:
[1.0, 1.0]
These are 1 to k-nearest neighbor's distances. We need to find their indices, a simple list.index function will only return the first match, so we will write the all_indices function in order to retrieve all the indices:
def all_indices(value, inlist):
out_indices = []
idx = -1
while True:
try:
idx = inlist.index(value, idx+1)
out_indices.append(idx)
except ValueError:
break
return out_indicesWith a value and list, all_indices will return all the indices where the value occurs in the list. We must convert our k smallest to a set:
ksmallest_set = set(ksmallest)
So, [1.0,1.0] becomes a set ([1.0]). Now, using a for loop, we can find all the indices of the elements:
# Get the indices of the K smallest elements for x in ksmallest_set: ksmallest_idx.append(all_indices(x,distances))
We get two indices for 1.0; they are 1 and 2:
ksmallest_idx = [item for sublist in ksmallest_idx for item in sublist]
The next for loop is to convert a list of the lists to a list. The all_indices function returns a list, and we then append this list to the ksmallest_idx list. Hence, we flatten it using the next for loop.
Finally, we add the k smallest neighborhood to our dictionary:
k_distance_neig[i].extend(zip(ksmallest,ksmallest_idx))
We then add tuples where the first item in the tuple is the distance and the second item is the index of the nearest neighbor. Let's print the k-distance neighborhood dictionary:
defaultdict(<type 'list'>, {0: [(1.0, 1), (1.0, 3)], 1: [(1.0, 0), (1.0, 2)], 2: [(1.0, 1), (1.0, 3)], 3: [(1.0, 0), (1.0, 2)], 4: [(4.0, 3), (5.0, 0)]})In step 4, we calculate the LRD. The LRD is calculated using the reachability distance. Let's recap both the definitions:
- The reachability distance from P to Q is defined as the maximum of the distance between P and its kth nearest neighbor, and the distance between P and Q. The following notation may help clarify this:
Reachability distance (P ß Q) = > maximum(K-Distance(P), Distance(P,Q))
- The Local Reachability density of P (LRD(P)) is the ratio of the k-distance neighborhood of P and the sum of the reachability distance of k and its neighborhood:
#Local reachable density local_reach_density = defaultdict(float)
We will first declare a dictionary in order to store the LRD:
for i in range(instances.shape[0]): # LRDs numerator, number of K distance neighbourhood no_neighbours = len(k_distance_neig[i]) denom_sum = 0 # Reachability distance sum for neigh in k_distance_neig[i]: # maximum(K-Distance(P), Distance(P,Q)) denom_sum+=max(k_distance[neigh[1]][0],neigh[0]) local_reach_density[i] = no_neighbours/(1.0*denom_sum)
For every point, we will first find the k-distance neighborhood of that point. For example, for i = 0, the numerator would be len (k_distance_neig[0]), 2.
Now, in the inner for loop, we calculate the denominator. We then calculate the reachability distance for each k-distance neighborhood point. The ratio is stored in the local_reach_density dictionary.
Finally, in step 5, we calculate the LOF for each point:
for i in range(instances.shape[0]): lrd_sum = 0 rdist_sum = 0 for neigh in k_distance_neig[i]: lrd_sum+=local_reach_density[neigh[1]] rdist_sum+=max(k_distance[neigh[1]][0],neigh[0]) lof_list.append((i,lrd_sum*rdist_sum))
For each data point, we calculate the LRD sum of its neighbor and the reachability distance sum with its neighbor, and multiply them to get the LOF.
The point with a very high LOF is considered an outlier. Let's print lof_list:
[(0, 4.0), (1, 4.0), (2, 4.0), (3, 4.0), (4, 18.0)]
As you can see, the last point has a very high LOF compared with the others and hence, it's an outlier.
