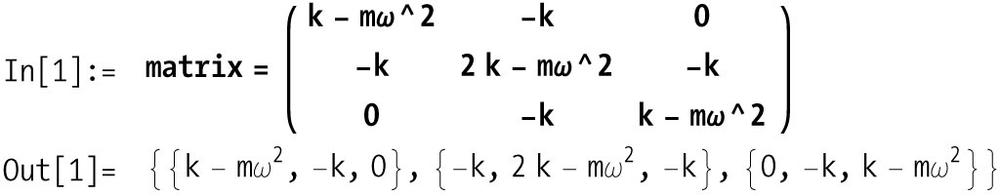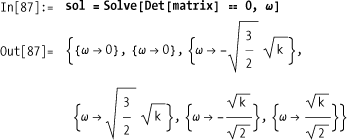Mmm—but it’s poetry in motion And when she turned her eyes to me As deep as any ocean As sweet as any harmony Mmm—but she blinded me with science And failed me in geometry
When she’s dancing next to me "Blinding me with science—science!" "Science!"
I can hear machinery "Blinding me with science—science!" "Science!"
—Thomas Dolby, "She Blinded Me With Science"
Scientists and engineers make up a large part of the Mathematica user base, and it is hard to think of any scientific or engineering practitioner, no matter how specialized, who could not benefit from Mathematica. I am neither a scientist nor an engineer by profession, but just fiddling around with Mathematica has given me insights into scientific and engineering ideas that otherwise would have taken many years of study.
The goals of this chapter are threefold. First, I want to illustrate techniques for organizing solutions to problems. Many science and engineering problems require numerous variables, and organization becomes paramount. There is not one correct way to organize complex solutions, but I provide two different approaches in 13.6 Solving Basic Rigid Bodies Problems and 13.11 Modeling Truss Structures Using the Finite Element Method. The second goal is to take some of the theoretical recipes covered in earlier chapters and apply them to real-world problems. I often see posts on Mathematica’s primary mailing list questioning the usefulness of function or pattern-based programming on real-world problems. Other posters express a wish to use these techniques but can’t get themselves on the right track. This chapter contains recipes to which most scientists and engineers can relate, and all use a mixture of functional and pattern-based ideas. An auxiliary goal is to take some of the functions introduced in Chapter 11 and make each the focus of a recipe. The third goal of the chapter is to introduce some special features of Mathematica that we did not have occasion to discuss in earlier recipes.
One important feature introduced in Mathematica 6 that gained
momentum in version 7 is curated data sources. These high-quality data
sources alone are worth the cost of admission to Mathematica’s user
base. 13.1 Working with Element Data through 13.4 Working with Genetic Data and Protein Data discuss some
sources pertinent to the sciences. Chapter 14 includes recipes related to financial
data sources. All these sources have a uniform, self-describing
structure. You can query any data source for the kinds of data it
provides using syntax DataSource["Properties"]. This will give you a
list of properties. Each property describes an important subset of the
data held by the source. You use the properties along with keys to
retrieve particular values. For example, ElementData[1, "AtomicWeight"] gives 1.00794,
the atomic weight of hydrogen. Once you master the data source concept,
you will quickly be able to leverage new data sources as they become
available.
13.5 Modeling Predator-Prey Dynamics applies the
discrete calculus function RSolve
from 11.10 Solving Difference Equations to solve a standard
predator-prey problem. Here I also demonstrate how Mathematica’s
interactive features can be used to explore the solution space and gain
insight into the dynamics of the problem.
In 13.6 Solving Basic Rigid Bodies Problems, I solve
a relatively straightforward problem in rigid body dynamics. The primary
purpose of this recipe is to illustrate one way you might organize a
problem with many objects and many parameters per object. This recipe
highlights Mathematica’s flexible ways of creating names of things, an
ability you should exploit when modeling complex problems. 13.11 Modeling Truss Structures Using the Finite Element
Method uses the topic of
finite element method (FEM) to illustrate an alternate way to organize a
problem that uses a lot of data and a variety of related functions. The
interface developed here follows a trend that is becoming more popular
in new Mathematica features (e.g., LinearModelFit).
13.7 Solving Problems in Kinematics through 13.10 Modeling Electrical Circuits focus on applied differential
equations. Here I solve some problems symbolically using DSolve and some problems numerically using
NDSolve. These recipes show how to
set up initial and boundary conditions, how to leverage Fourier series
in obtaining solutions, and how to visualize solutions.
You want to perform computations that take as input information about the chemical elements. You may also want to create visual displays of this information for reference or classroom use.
You can list the names of all the elements using ElementData[] or the name of the
nth element using ElementData[n].
In[1]:= ElementData[] Out[1]= {Hydrogen, Helium, Lithium, Beryllium, Boron, Carbon, Nitrogen, Oxygen, Fluorine, Neon, Sodium, Magnesium, Aluminum, Silicon, Phosphorus, Sulfur, Chlorine, Argon, Potassium, Calcium, Scandium, Titanium, Vanadium, Chromium, Manganese, Iron, Cobalt, Nickel, Copper, Zinc, Gallium, Germanium, Arsenic, Selenium, Bromine, Krypton, Rubidium, Strontium, Yttrium, Zirconium, Niobium, Molybdenum, Technetium, Ruthenium, Rhodium, Palladium, Silver, Cadmium, Indium, Tin, Antimony, Tellurium, Iodine, Xenon, Cesium, Barium, Lanthanum, Cerium, Praseodymium, Neodymium, Promethium, Samarium, Europium, Gadolinium, Terbium, Dysprosium, Holmium, Erbium, Thulium, Ytterbium, Lutetium, Hafnium, Tantalum, Tungsten, Rhenium, Osmium, Iridium, Platinum, Gold, Mercury, Thallium, Lead, Bismuth, Polonium, Astatine, Radon, Francium, Radium, Actinium, Thorium, Protactinium, Uranium, Neptunium, Plutonium, Americium, Curium, Berkelium, Californium, Einsteinium, Fermium, Mendelevium, Nobelium, Lawrencium, Rutherfordium, Dubnium, Seaborgium, Bohrium, Hassium, Meitnerium, Darmstadtium, Roentgenium, Ununbium, Ununtrium, Ununquadium, Ununpentium, Ununhexium, Ununseptium, Ununoctium} In[2]:= ElementData[1] Out[2]= Hydrogen
Mathematica will return properties of an element if given its number and the name of the property.
In[3]:= Row[{ElementData[1], ElementData[1, "AtomicNumber"], ElementData[1, "AtomicWeight"], ElementData[1, "Phase"]}, " "] Out[3]= Hydrogen 1 1.00794 Gas
You can see from the list of all properties that
Mathematica has a comprehensive database of elemental data. Be aware
that CommonCompoundNames will pull
in a lot of data if you use it with a common element like
hydrogen.
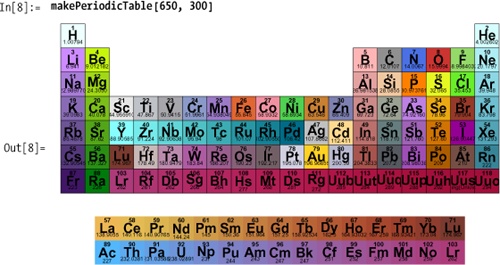
The most obvious application of ElementData is to create a periodic table.
The ElementData documentation shows
code for a simple table. Here I show a more ambitious one, complete
with Tooltip.
You want to perform computations that take as input information about the chemical compounds. You may also want to create visual displays of this information for reference or classroom use.
ChemicalData is a curated
data source. You can request chemical information by common names,
registry numbers, IUPAC-like names, or structure strings.
The list of properties of chemical compounds is quite impressive. The table below lists a random subset of the full list of 101 properties.
In[14]:= Partition[Sort[RandomSample[ChemicalData["Properties"], 30]], 3] //
TableForm
Out[14]//TableForm=
AcidityConstant BoilingPoint CHStructureDiagram
CIDNumber CompoundFormulaDisplay CriticalPressure
CriticalTemperature FlashPointFahrenheit FormattedName
HildebrandSolubility IUPACName MDLNumber
MeltingPoint NFPAHazards NFPAHealthRating
NFPALabel NonStandardIsotopeCount NonStandardIsotopeNumbers
PartitionCoefficient Phase Resistivity
RotatableBondCount SideChainAcidityConstant SpaceFillingMoleculePlot
StructureDiagram TautomerCount TopologicalPolarSurfaceArea
VaporPressureTorr VertexTypes ViscosityAt the time of this writing, Mathematica has curated data on over 34,300 compounds, subdivided into 67 classes.
In[15]:= Length[ChemicalData[]] Out[15]= 34336 In[16]:= ChemicalData["Classes"] Out[16]= {AcidAnhydrides, AcidHalides, Acids, Alcohols, Aldehydes, Alkanes, Alkenes, Alkynes, Alloys, Amides, Amines, AminoAcidDerivatives, AminoAcids, Arenes, Aromatic, Bases, Brominated, Carbohydrates, CarboxylicAcids, Catalysts, Cations, Ceramics, Chiral, Chlorinated, Dendrimers, Esters, Ethers, Fluorinated, Furans, Gases, Halogenated, HeavyMolecules, Heterocyclic, Hydrides, Hydrocarbons, Imidazoles, Indoles, Inorganic, Iodinated, IonicLiquids, Ketones, Ligands, Lipids, Liquids, MetalCarbonyls, Monomers, Nanomaterials, Nitriles, Organic, Organometallic, Oxides, Phenols, Piperazines, Piperidines, Polymers, Pyrazoles, Pyridines, Pyrimidines, Quinolines, Salts, Solids, Solvents, Sulfides, SyntheticElements, Thiazoles, Thiols, Thiophenes}
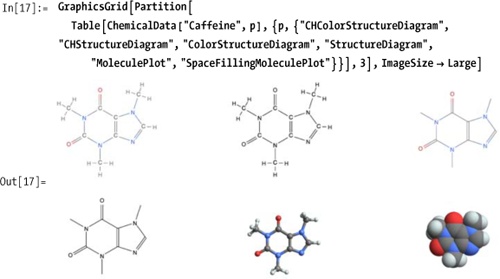
There are six kinds of structural diagrams that can be used to visualize these compounds. Here, for example, are representations for what may be one of your favorites, for better or worse—caffeine.
You can use the data to analyze relationships between
properties. Here I show a plot of inverse vapor pressure to boiling
point for all liquids with a Tooltip around each point so outliers are
easy to identify. Cases is used to
filter out any MissingData
entries.
You want to perform computations that take as input information about the elementary particles. You may also want to create visual displays of this information for reference or classroom use.
In[19]:= ParticleData["Classes"]
Out[19]= {Baryon, BBBarMeson, Boson, BottomBaryon, BottomMeson,
CCBarMeson, CharmedBaryon, CharmedMeson, Fermion, GaugeBoson,
Hadron, Lepton, LongLived, Meson, Neutrino, Pentaquark, Quark,
Stable, StrangeBaryon, StrangeCharmedBaryon, StrangeCharmedMeson,
StrangeMeson, UnflavoredBaryon, UnflavoredMeson}It is easy to create functions that generate tables of particle
information. The function particleTable accepts a list of one or more
class memberships (e.g., Baryon,
LongLived, and others from ParticleData["Classes"]) and a list of
properties to use as columns. The helper function particleData reformats "QuarkContent" into a more concise
representation. You will often want to filter out entries that are
missing since there is only partial data available for exotic
particles.
Create a table of long-lived baryons. A baryon is a particle made of three quarks, and long-lived refers to particles whose lifetime is greater than 10–20 seconds.
The list of properties available in particle data are as follows:
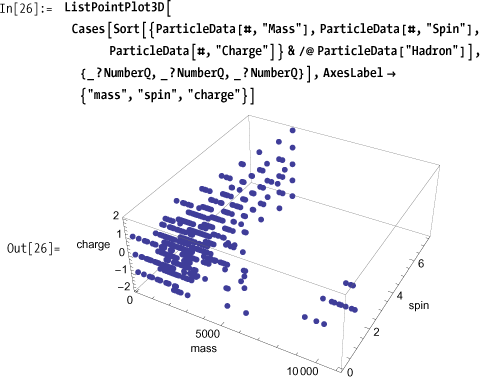
A scatter plot of mass versus spin versus charge shows large voids where there are no known particles (or where the values are unknown).
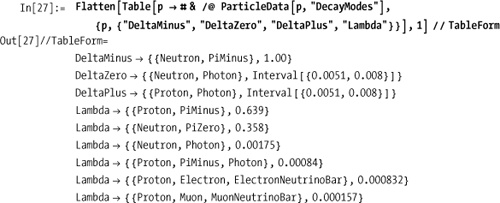
DecayModes and
FullDecayModes list the ways the
particle can decay; FullDecayModes
also lists those predicted by theory but not observed in detectors.
The number (or interval) display with the decay mode is the branch
ratio.
You want to use Mathematica’s pattern matching and computational
capabilities to develop bioinformatics applications. GenomeData and ProteinData provide the raw materials for
this application.
Get the first 100 nucleobases (or, simply, bases) on the male X chromosome.
Get the first 10 proteins known to Mathematica and show number of amino acids in its sequence.
In[29]:= {#, ProteinData [#, "SequenceLength"]} & /@ Take[ProteinData[], 10] // TableForm Out[29]//TableForm= A1BG 495 A2M 1474 NAT1 290 NAT2 290 SERPINA3 423 AADAC 399 AAMP 434 AANAT 207 AARS 968 ABAT 500
Find five other chromosomes that have sequences that match the first 50 bases of chromosome-1 in the human genome. Strands of the chromosome are indicated as 1 or -1.
In[30]:= GenomeLookup[GenomeData[{"Chromosome1", {1,50}}], 5]
Out[30]= {{{Chromosome1, 1}, {1, 50}}, {{Chromosome1, 1}, {7, 56}},
{{Chromosome1, 1}, {13, 62}}, {{Chromosome3, -1}, {116621, 116670}},
{{Chromosome3, -1}, {116615, 116664}}}At the time of writing, Mathematica has data on 27,479 proteins and 39,920 genes.
In[31]:= {Length[ProteinData[]], Length[GenomeData[]]}
Out[31]= {27479, 39 920}The following is a list of properties of the proteins. This data
is somewhat incomplete: some of the values are not known or have not
been updated in Wolfram’s database. The good news is that it improves
over time, so there is likely more data when you’re reading this than
when I wrote it. Notice how this sample is ordered in columns, whereas
prior recipes showed similar lists in rows. All you need is Transpose and a bit of math to get the
desired number of columns.
In[32]:= Module[{props = ProteinData["Properties"]}, With[{nCols = Ceiling[Length[props]/3]}, Transpose[Partition[props, nCols, nCols, 1, "" ]]]]//TableForm Out[32]//TableForm= AdditionalAtomPositions DNACodingSequence MolecularWeight AdditionalAtomTypes DNACodingSequenceLength MoleculePlot AtomPositions DomainIDs Name AtomRoles DomainPositions NCBIAccessions AtomTypes Domains PDBIDList BiologicalProcesses Gene PrimaryPDBID CellularComponents GeneID SecondaryStructureRules ChainLabels GyrationRadius Sequence ChainSequences Memberships SequenceLength DihedralAngles MolecularFunctions StandardName
One property that is sparsely populated is MoleculePlot. At the time of writing, the
only protein beginning with "ATP" that has a MolecularPlot is ATP7BIsoformA.
GenomeData likewise contains
a wealth of information. Here I show the properties available.
In[34]:= Module[{props = GenomeData["Properties"]}, With[{nCols = Ceiling[Length[props]/3]}, Transpose[Partition[props, nCols, nCols, 1, ""]]]]//TableForm Out[34]//TableForm= AlternateNames GBandStainingLevels Orientation AlternateStandardNames GenBankIndices ProteinGenBankIndices BiologicalProcesses GeneID ProteinNames CellularComponents GeneOntologyIDs ProteinNCBIAccessions Chromosome GeneType ProteinStandardNames CodingSequenceLists InteractingGenes PubMedIDs CodingSequencePositions IntronSequences SequenceLength CodingSequences LocusList StandardName ExonSequences LocusString TranscriptGenBankIndices FullSequence Memberships TranscriptNCBIAccessions FullSequencePosition MIMNumbers UniProtAccessions GBandLocusStrings MolecularFunctions UnsequencedPositions GBandScaledPositions Name UTRSequences GBandStainingCodes NCBIAccessions In[35]:= GenomeData["ACOT9", "ProteinNames"] Out[35]= {acyl-Coenzyme A thioesterase 2, mitochondrial isoform a, acyl-Coenzyme A thioesterase 2, mitochondrial isoform b} In[36]:= GenomeData["ACOT9", "Memberships"] Out[36]= {ChromosomeXGenes, Genes, Hydrolase, Mitochondrion, ProteinBinding, ProteinCoding} In[37]:= GenomeData["ACOT9", "CellularComponents"] Out[37]= {Mitochondrion} In[38]:= GenomeData["ACOT9", "MolecularFunctions"] Out[38]= {AcetylCoAHydrolaseActivity, CarboxylesteraseActivity, HydrolaseActivity, ProteinBinding}
You want to model a dynamic system consisting of populations of predators and prey to see how population levels evolve over time.
Consider a population of rabbits (prey) and foxes (predators)
with a specific growth rate for rabbits G and carrying capacity of their environment
K. The population dynamics can be
modeled by a pair of difference equations. See the Discussion section for more insight into the form
of these equations and the meaning of the constants.
NestList presents one
possible solution for deriving the dynamics of the population over
time from an initial starting point.
This shows the rabbit population doing what rabbits do for many generations as the fox population slowly increases due to the increasing food supply. An inflection point is reached, and the fox population begins to take off with a resulting collapse in the rabbit population. Eventually the system reaches equilibrium.
The equation for rabbits assumes that rabbits follow the logistic model of exponential growth limited by the carrying capacity of the environment and then subtracts a term proportional to the number of rabbits and foxes where the constant 0.0001 reflects the efficiency of the predators. The equation for foxes assumes that the fox population is proportional to the ability to catch rabbits (same term from first equation) minus some natural death rate (here 2 percent of the population).
NestList provides a very
simple solution to this model, but it is not the best choice if, due
to efficiency, you want to create an interactive model using Manipulate. Luckily, Mathematica 7 has new
capabilities for discrete math that provide an alternate solution
path. RecurrenceTable is a new
function that will generate the list of solutions of specified length
given a recurrence relation.
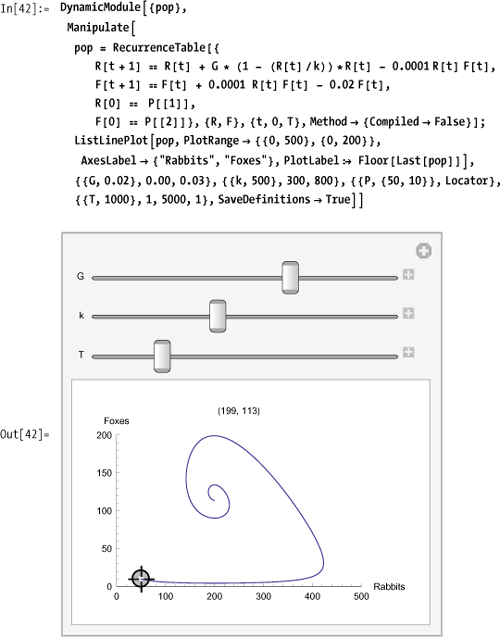
This interactive model allows you to position the locator at the initial population levels for rabbits and foxes and allows you to adjust the growth rate, carrying capacity, and number of iterations. The plot title displays the end value of rabbits and foxes.
More elaborate predator-prey models can be found at the Wolfram Demonstration Project: http://bit.ly/mUVGS and http://bit.ly/21GfLm.
You want to compute mass, center of mass, and moment of inertia as a prerequisite to solving dynamical problems involving rigid bodies.
The basic equation for computing the center of mass given a collection of discrete point masses is
where cmi is the center of mass of each point and mi is its mass. The numerator of this equation is called the first moment. Another name for the center of mass is the centroid.
In[43]:= centerMass[particles_] := Module[{totalMass, firstMoment}, {firstMoment, totalMass } = Sum[{mass @ particles[[i]] centroid @ particles[[i]], mass @ particles[[i]]}, {i, Length[particles]}]; firstMoment/ totalMass] In[44]:= mass@car = 1000.; centroid @ car = {100, 100}; mass @ driver = 86.; centroid@driver = {103, 101} ; mass@fuel = 14.2; centroid @ fuel = {93, 100}; centerMass[{car, driver, fuel}] Out[50]= {100.144, 100.078}
The solution is fairly elementary from a physical point of view,
but it may look a bit mysterious from a Mathematica coding point of
view. The solution is coded using Mathematica’s prefix notion. Recall
that f@x is prefix notation for
f[x]. This notation is appealing
for modeling problems because it is concise and readable (simply
replace the @ with "of" as you read
the code). Notice how I use the same notation with the problem objects
car, driver, and fuel.
Now suppose this notation does not appeal to you;
perhaps you like to model the physical objects as lists or some other
notation like object[{mass,centroid}]. Does this mean you
need to reimplement the centerMass
function? Not at all. Simply define the function’s mass and centroid
for your preferred representation, and you are all set.
In[51]:= mass[object[{m_,__}]] := m In[52]:= centroid[object[{_, c_,__}]] := c In[53]:= centerMass[{object[{1000, {100, 100}}], object[{86, {103, 101}}], object[{14.2, {93, 100}}]}] Out[53]= {100.144, 100.078}
Another important property of rigid bodies is the mass moment of
inertia about an axis. These values are important when solving
problems involving rotation of the body. The general equation for the
mass moment of inertia involves integration over infinitesimal point
masses that make up the body, but in practice problems, equations for
known geometries are typically used. One way to approach this in
Mathematica is to use a property called shape and rely on pattern matching to select
the appropriate formula. Each of these functions returns a list in the
form {Ixx, Iyy, Izz}, giving the
moment of inertia about the x-, y-, and z-axis, respectively.
In[1]:= massMomentOfInertia[o_] /; shape@o == "circularCylinder" := Module[{i1, i2}, i1 = ((mass@o radius@o^2)/4) + ((mass@o length @o ^2)/12); i2 = ((mass@o radius@o^2)/2); {i1, i1, i2} ] massMomentOfInertia[o_] /; shape@o == "circularCylindricalShell" := Module[{i1, i2}, i1 = ((mass@o radius@o^2)/2) + ((mass@o length @o ^2)/12); i2 = ((mass@o radius@oA2)); {i1, i1, i2} ] massMomentOfInertia[o_] /; shape@o == "rectangularCylinder" := Module [{ixx, iyy, izz}, ixx = ((mass@o (height@o + length@o) ^2)/12); iyy = ((mass@o (width@o + length@o) ^2)/12); izz = ((mass@o (width@o + height@o) ^2)/12); {ixx, iyy, izz} ] massMomentOfInertia[o_] /; shape@o == "sphere" := Module[{i}, i = (mass@o (2 radius@o ^2)/5); {i, i, i} ] massMomentOfInertia[o_] /; shape@o == "sphericalShell" := Module[{i}, i = (mass@o (2 radius@o A2) /3); {i, i, i} ] In[59]:= shape@car = "rectangularCylinder"; length@car = 4.73; width@car = 1.83; height@car = 1.25; In[63]:= massMomentOfInertia[car] Out[63]= {2980.03, 3586.13, 790.533} In[64]:= shape@car = "circularCylindricalShell"; radius@car = 1.83; In[66]:= massMomentOfInertia[car] Out[66]= {3538.86, 3538.86, 3348.9}
You want to demonstrate standard problems in kinematics, like those you typically find in first-year physics studies.
The basic equations of kinematics are as follows.
In[67]:= acceleration1[deltaT_, v1_, v2_] := (v2 - v1) / deltaT acceleration2[deltaT_, v1_, deltaS_] := 2 (deltaS - v1 deltaT) / (deltaT^2) acceleration3[v1_, v2_, deltaS_] := (v2^2 - v1^2) / (2 deltaS) distance[a_, v1_, deltaT] := (adeltaT^2 / 2) + v1 deltaT distance1[a_, v1_, v2_] := (v2^2 - v1^2) / (2 a) distance2[deltaT, v1_, v2_] := (deltaT / 2) (v1 + v2) time1[a_, v1_, v2_] := (v2 - v1) / a time2[a_, v1_, deltaS] := (Sqrt[v1^2 + 2 + 2adeltaS] - v1) / a time3[v1_, v2_, deltaS] := (2 deltaS) / (v1 + v2) velocity1[a_, v2_, deltaT_] := v2 - a deltaT velocity2[a_, deltaS_, deltaT_] := (deltaS/ deltaT) - (a deltaT/ 2) velocity3[a_, v2_, deltaS_] := Sqrt[v2^2 - 2 a deltaS]
Given these equations, you can solve a variety of problems. For example, how far will a bullet drop if shot horizontally from a rifle at a target 500 m away if the initial velocity is 800 m/s? Ignore drag, wind, and other factors.
First, compute how long the bullet remains in flight before hitting the target by taking the initial and final velocity to be the same.
In[79]:= timeTraveled = time3[800, 800, 500] // N
Out[79]= 0.625Given the acceleration due to gravity is 9.8 m/s2, compute the distance dropped by setting the initial vertical velocity component to zero.
In[80]:= distanceDropped = distance[9.8, 0, timeTraveled]
Out[80]= 1.91406The bullet drops almost 2 meters.
The solution works out a simple problem by working first in the x direction and then plugging the results into an equation in the y direction. In more complex problems, it is often necessary to use vectors to capture the velocity components in the x, y, and z directions. Consider a game or simulation involving a movable cannon and a movable target of varying size.
Imagine the cannon is fixed to the side of a fortress such that
the vertical height (z direction in this example)
is variable but the x and y
position is fixed. The length, angle of elevation (alpha), left-right
angle (gamma), and muzzle velocities are also variable. You require a
function that gives the locus of points traversed by the shell given
the cannon settings and the time of flight. Here we use Select to filter the points above ground
level (positive in the z direction). The function
returns a list of values of the form {{x1,y1,z1,t1}, ..., {xn,yn,zn,tn}}, where
each entry is the position of the shell at the specified time.
Chop is used only to replace
numbers close to zero by zero. Note that in each dimension, the basic
kinematic equations are in play, but since the inputs are in terms of
angles, some basic trigonometry is needed to get the separate
x, y, and z components.
Velocity is constant in the x-y plane (we are still ignoring drag),
and the z-axis uses the initial velocity component and the fall of the
shell due to gravity.
You can also create a function that computes the instantaneous velocity components at a specified time.
In[82]:= velocity[velocity_, alpha_, gamma_, t_] := With[{g = 9.8}, Chop[ If[t > 0, {velocity * Cos[alpha * Pi / 2], velocity * Cos[gamma * Pi], velocity Sin[alpha * Pi/ 2] - g t}, {0., 0., 0.}] ] ]
Since the plan is to create a simulation, you need a function that figures out when the shell intersects with the target. For simplicity, assume the shape of the target is a box.
You can set the simulation up inside of a Manipulate so that you can play around with
all the variables.
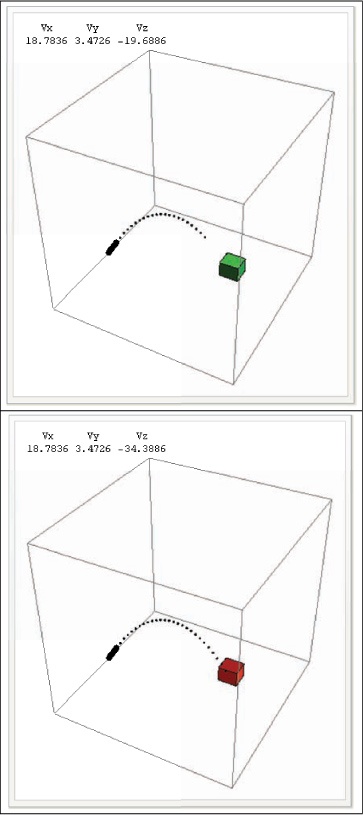
The initial output of the Manipulate is shown in Out[85] above. The
path of the bullet is displayed up until the point in time specified
by the time control, so the box turns red after it is hit by a shell.
The instantaneous velocity of the shell is displayed for the current
value of time. The Vz will be
negative when the shell is falling. Figure 13-1 shows two frames
from the Manipulate, at a time
before impact and a time after.
David M. Bourg’s Physics for Game Developers (O’Reilly) has an example of the cannon problem where wind drag is introduced. Keep in mind that the author uses the y-axis as the vertical whereas the code in this recipe uses the z-axis.
Mathematical Methods Using Mathematica by Sadri Hassani (Springer) has solutions to similar problems using differential equations which consider drag, curvature of the earth, and nonconstant acceleration at large distances from the earth’s surface (see Chapter 6).
You want to compute the normal modes for a system of
identical masses connected by identical springs. Normal modes are
natural or resonant frequencies of the entire system. The system this
recipe considers consists of n>1
masses connected by n-1 springs on
a frictionless surface. Figure 13-2 shows an
example for n=3.
Here I state, without proof (refer to See Also section), that these systems take the
form of n simultaneous linear equations whose
matrix representation is tridiagonal. That is a matrix with nonzero
entries along the main diagonal and adjacent minor diagonals and zero
entries in all other elements. The corner entries of the main diagonal
are special since they represent masses that are free on one side and
take the form k - m*ω^2, where k is
the spring constant, m is the mass,
and ω is the angular frequency. The off corner entries represent the
masses with springs on both sides and take the form 2*k - m*ω^2. The minor diagonals are all -k. Here
I solve the three mass problems, and in the discussion, I show how to
create a general solver for the n mass case.
Nontrivial solutions to this system leave the matrix as
noninvertible; hence, the determinant is zero. Use Solve to find the frequencies in terms of
k.
You don’t care about the solutions with negative or zero frequencies, so you can filter these out to obtain two physically interesting resonant frequencies.
Given the frequencies, you can solve the system to get the
amplitudes. The first solution gives al ==
a3 and a2 == -2a1, with
the alternative of k == 0 being
physically uninteresting. This solution has the outer masses moving in
unison in the same direction while the inner mass compensates by
moving in the opposite direction with twice the amplitude.
In[89]:= Reduce[Dot[(matrix /. sol[[1]]), {a1,a2,a3}] == 0, {a1,a2,a3}]
Out[89]= (a2 ==-2a1&&a3 = al) || k = 0The second solution gives a2 ==
0 and a3 == -al with the
alternative of k == 0 being
physically uninteresting. This is a solution with the center mass at
rest and the outer masses moving toward and then away from the
center.
In[90]:= Reduce[Dot[(matrix /. sol[[2]]), {a1,a2,a3}] == 0, {a1,a2,a3}]
Out[90]= (a2 = 0&&a3 == -al) || k = 0To solve the general n-mass system, we need
a way to synthesize a tridiagonal matrix of the proper form. For this,
SparseArray and Band are just what the doctor ordered. When
using sparse matrix, rules that come earlier override rules that come
later. This works to your favor because it allows the case where
n == 2 to be handled without any
conditional logic stemming from the fact that there are no 2*k - m*ω^2 terms when n == 2.
For the general solution, you want to use NSolve with specific values of m and k
because roots of polynomials with degree greater than five are likely
to give Solve trouble. Here I solve
a 10-mass system with k == 1 and
m == 1. Chop is used to remove
residual imaginary values and Cases
filters out zero and negative solutions because they are physically
uninteresting.
You can find derivations of the systems solved in this recipe in many advanced physics and linear algebra books. In particular, Mathematical Methods Using Mathematica by Sadri Hassani provides a nice mix of practical physics and Mathematica techniques, although the most recent edition is written for versions of Mathematica prior to 6 and therefore does not always indicate the best technique to use for current versions.
You want to model the dynamics of a vibrating string after it is released from a particular deformation.
This solution is a particular solution to the one-dimensional
wave equation D[u[x,t], {x,2}] == c^2
D[u[x,t],{t,2}] where u[x,t] gives the position of the string at
point x and time t. The
general solution to the wave equation can be obtained using
DSolve.
The general solution is not very helpful because it is specified
in terms of two unknown functions, C[1] and C[2]. In theory, you could specify boundary
conditions and initial conditions, but DSolve is very limited in its ability to
find solutions to partial differential equations. This problem is
better handled numerically with NDSolve.
First we need a specification for the shape of the string at
t = 0. For simplicity, I’ll use the Sin function that will give a width of
L units. Here I use Plot to show the initial defection of the
string.
To use NDSolve to model the
vibrating string, you must provide initial and boundary conditions.
The initial condition states that u[0,x] =
string[x]. In other words, at the start, the string has the
position depicted previously. You must also specify the initial
velocity of the string, which is the first derivative with respect to
time. The obvious choice for initial velocity is zero. Using input
form, this would be entered as Derivative[1,
0][u][0, x] == 0. This operator notation was explained in
11.4 Differentiating Functions. The two boundary
conditions specify that the ends of the string are anchored at
position 0 and L, u[t, 0] == 0, and u[t, L]
== 0.
Although DSolve can
deal with some partial differential equations (PDEs), it is limited in
its ability to derive specific solutions given initial and boundary
conditions. Therefore, it is better to use NDSolve on PDEs, as I’ve done in the
solution. However, it is not difficult to pose problems that NDSolve will have a hard time with and
ultimately fail to solve. Consider trying to solve the wave equation
with an initial position that contains a discontinuity.
If you try to use string2 in
the solution shown in In[99] above, it will likely run for a very long
time, consuming memory and finally failing. However, this situation is
not entirely hopeless. One technique is to produce an approximation to
string2 using Fourier series. Using
Fourier series, I obtained the following Sin expansion, called sinString2:
In[102]:= sinString2[x_] = 0.285325252629769` Sin[(Pi*x)/5] - 0.033193742967516204` Sin[(3*Pi*x)/5] + 0.013117204588138661` Sin[πx] - 0.007723288156504195` Sin[(7*Pi*x)/5] + 0.005695145921372713` Sin[(9*Pi*x)/5] - 0.004945365736699312` Sin[(11*Pi*x)/5];
Below I plot both functions to demonstrate how closely sinString2 approximates string2 while smoothing out the
discontinuity at the apex.
There is an exact solution to the triangular wave, although it
isn’t derived here (refer to the See Also
section). It is given by this infinite sum, which Mathematica can
solve using a special function LerchPhi. This solution is too complex to
use in an animation, but you can use it to verify that the approximate
solution is quite good.
Plotting a few snapshots of the exact solution over time tells us that the approximate solution is more than adequate and, in some sense, superior because it is far less computationally intense.
There are many ways to approach the solution to the wave equation. When this problem is solved by hand, separation of variables is often employed. See Advanced Engineering Mathematics by Erwin Kreyszig (John Wiley) for a step-by-step example. Warning: this book is not a Mathematica reference, but the problems are worked out in enough detail that you can easily see your way to creating your own Mathematica-based solutions.
You want to understand how electrical circuits consisting of resistors, capacitors, and inductors behave.
The differential equation governing an RLC circuit is L I'' + R I' + I/C = E(t), where I is current, L is inductance, R is resistance, C is capacitance, and E(t) is the electromotive force (commonly
known as voltage). Modeling the system means understanding how the
current varies as you drive the system with a particular timing
varying voltage. Let’s consider a common sinusoidal voltage and solve
the system assuming that the charge and current are zero at t=0. Setting the problem up with the context
of a With allows you to solve the
problem for different values of inductance, capacitance, resistance,
frequency, and voltage.
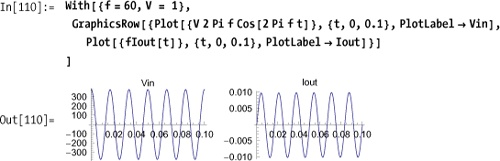
By plotting the input voltage and output current, you can see they have the same basic shape and frequency except for a phase shift.
A more interesting example uses a nonsinusoidal wave,
such as a triangular wave. Conveniently, Mathematica 7 has a function
TriangleWave[t] that suits our
purpose.
However, the discontinuities in this waveform throw DSolve for a loop. To work around this,
represent the triangular wave by its Fourier series. This will give a
very close approximation without the discontinuities at the extremes.
This will allow you to use DSolve.
Notice how the RLC circuit responds to the triangular
wave input by smoothing the current flow to an approximately
sinusoidal form. As an exercise, you can try this same example using
SquareWave, SawtoothWave, or other
functions of your own design.
You want to build a model based on the finite element method (FEM). You want to organize the model in a manner that allows you to obtain the solution as well as other intermediate results and structural diagrams.
The FEM has a wide range of engineering applications. In this recipe, I will limit the discussion to structures composed of linear elements known as trusses. See the figure shown in the "Discussion" section. Here my focus will be on the organization of the solution within Mathematica rather than on the underlying theory. Therefore, all results will be present without derivation of the underlying mathematics. Please refer to the references in the See Also section.
To begin, you will need a means to represent the elements. I use
a structure called linearElement
that specifies two endpoints called nodes ({{x1, y1}, {x2, y2}}), an area, and a measure of stiffness called
Young’s Modulus (YM).
linearElement[{{x1, y1}, {x2, y2}}, area, YM]In addition, you need a means for specifying the x and y components of the force at each node.
force[{x, y}, fx, fy]Furthermore, at each node there is a computed displacement in the x and y direction. The FEM literature uses the variable u for x displacements and v for y displacements Typically, each node is sequentially numbered, so you would have ul, vl, u2, v2, and so on. I will not use a sequential numbering here, bocuuso ouch node is uniquely identified by its coordinates, and given Mathematica’s liberal representation of variables, it is much more convenient to specify nodal displacements using coordinates.
u[x1, y1] (*The displacement in the x direction at node {x1,y1}*) v[x1, y1] (*The displacement in the y direction at node {x1,y1}*)
With these conventions established, I proceed by defining a series of helper functions that will be needed later. I provide a brief description of each function but, for brevity, defer more detail to the Discussion section.
Each element in the model is governed by a system of linear
equations. The system is naturally represented by a symmetric matrix.
The symmetry takes the form {{A,-A},{-A,A}} where A is a block matrix.
A location vector provides a means for locating the position of
the local element matrices computed by linearElementMatrix within a larger global
matrix that represents the system over all elements.
In[119]:= assemblyLocationVector[linearElement[{n1_, n2_},__], allnodes_] := Flatten[Position[allnodes, #]& /@ {u@@n1, v@@n1, u@@n2, v@@n2}]
This helper maps a node of the form {{x1,y1},{x2,y2}} to the corresponding force
components {{fx1,fy1},{{fx2,fy2}}}.
It does this by searching for the first match of the node within the
list of forces and transforming it to the desired form.
In[120]:= getExternalForces[{forcesforce}, node_] := Cases[{forces}, force[node, fu_, fv_] :> {fu, fv}, 1, 1]
This helper extracts the unique set of nodes from the
elements and places them in a canonical order, as defined by Union. This ordering is essential to the
construction of a consistent system of equations. See the Discussion section for details.
In[121]:= getNodes[{elementslinearElement}] := Union[{elements} /. linearElement[{n1_, n2_},__] :> Sequence[n1, n2]]
This helper is used to construct a replacement rule for forces.
Construct a global vector of all forces using a set of nodes in canonical order.
In[123]:= getForceVector[{forces__force}, nodes_] := Flatten[getExternalForces[{forces}, #]& /@ nodes]
Assemble the global matrix that defines the system of equations
over all elements using the local matrices for individual elements and
the location vectors that define the position of the local matrices
with the global matrix. Note that the global matrix is obtained by
summing the local matrices into the appropriate positions within the
global matrix. In other words, think of each member of locationVectors as specifying a submatrix
within the global matrix for which the corresponding member of
localMatrices is added.
In[124]:= assembleGlobalMatrix[localMatricies_, locationVectors_, numElements_, dimension_] := Module[{g}, g = Table[0, {dimension}, {dimension}] ; Do[g[[locationVectors[[i]], locationVectors[[i]] ]] += localMatricies[[i]], {i, 1, numElements}] ; g ]
A model consists of a collection of connected elements, the
external forces applied to the structure at one or more nodes, and the
boundary conditions that typically manifest as points where a node is
anchored, rendered immobile in the x, y, or both
directions. Here I organize a solution in the spirit of LinearModelFit covered in Chapter 12. That is, I construct an
object called a TrussModel, the
function of which is to organize the underlying data and then use that
object as the target for requests for certain properties relevant to
the FEM. As of Mathematica 6 and particularly in Mathematica 7, this
object-based methodology has emerged as a design pattern for
organizing solutions that involve large quantities of data or
collections of related functionality.
To proceed in this manner, you need a function for
creating the TrussModel and a
Format for displaying it. The
Format is syntactic sugar that
hides the details of the TrussModel, which could be quite
large.
In[125]:= createTrussModel[{elementslinearElement}, {forces___force}, boundaryNodes_] : = Module[{localMatrices, nodes, nodalVar, forceVec, locationVectors, degreesOfFreedom, globalMatrix, allForces, forceRules}, nodes = getNodes[{elements}]; nodalVar = Flatten[{u@@#, v@@#}&/@ nodes]; localMatrices = linearElementMatrix /@ {elements}; locationVectors = assemblyLocationVector [#, nodalVar] & /@ {elements}; globalMatrix = assembleGlobalMatrix[localMatrices, locationVectors, Length[{elements}], Length[nodalVar]]; degreesOfFreedom = Complement[Range[Length[nodalVar]], Flatten[Position[nodalVar, #]& /@ boundaryNodes]]; allForces = force [#, 0, 0]& /@ nodes; forceRules = makeForceRule /@ {forces}; allForces = allForces /. forceRules; forceVec = getForceVector[allForces, nodes]; TrussModel[{elements}, boundaryNodes, localMatrices, globalMatrix, nodalVar, forceVec, degreesOfFreedom, forces]] Format[TrussModel[elements_, boundaryNodes_, __ ]] := ToString[TrussModel[{Length[elements]}, {Length[boundaryNodes]}]]
The goal of a FEM analysis is to determine the behavior of the
structure from the behavior of the elements. For a system of trusses,
solve for the displacements at the joints, the axial forces, and axial
stresses. Following the proposed methodology, these will be accessed
as properties of the TrussModel.
The displacements property is
implemented as a functional pattern associated with the TrussModel. This notation may look somewhat
unusual but is quite natural from the standpoint of Mathematica’s
design. It simply states that when you see a pattern consisting of a
TrussModel and a literal argument,
"displacements", replace it with
the results of computing the displacements using data from the
TrussModel.
In[127]:= TrussModel[_, _, _, globalMatrix_, nodalVars_, forceVec_, degreesOfFreedom_, ___]["displacements"] := Flatten[Solve[Dot[globalMatrix[[degreesOfFreedom, degreesOfFreedom]], nodalVars[[degreesOfFreedom]]] == forceVec[[degreesOfFreedom]], nodalVars[[degreesOfFreedom]]]]
As a matter of convenience, you can make a property the
default property of the model by associating it with the invocation of
the model with no arguments. Of course, thus far I have defined only
one property, displacements, but it
was my intent to make this the default. In the discussion I derive
other properties of this model.
In[128]:= TrussModel[model][] := TrussModel[model]["displacements"]All this tedious preparation leads us to a solution that is very easy to use. Here is the TrussModel, depicted in Out[136] on Discussion. The example data is borrowed from a problem presented in Bhatti’s book (refer to the See Also section).
In[129]:= tm = createTrussModel[ {linearElement[{{0, 0}, {1500, 3500}}, 4000., 200*10^3], linearElement[{{1500, 3500}, {5000, 5000}}, 4000., 200*10^3], linearElement[{{0, 0}, {0, 5000}}, 3000., 200*10^3], linearElement[{{0, 5000}, {5000, 5000}}, 3000., 200*10^3], linearElement[{{0, 5000}, {1500, 3500}}, 2000., 70*10^3]}, {force[{1500, 3500}, 0, -150000]}, {u[0, 0], v[0, 0], u[5000, 5000], v[5000, 5000]}] Out[129]= TrussModel[{5}, {4}]
Now you can compute the nodal displacements at the nodes that are unsupported.
To complete the TrussModel,
we need to define more properties. It is nice to have a visual aid to
help diagnose problems in the setup of the model. A "diagram" property generates graphics. As
before, I need to develop some helper functions to take care of
certain tails. Each helper function has a placeholder for options
(opts___), but to keep the
implementation from getting any more complicated, I do not implement
any options. You could add options to control the level of detail, for
example, to include or suppress displacement arrows and labels. Other
options might be pass-through options to Graphics.
The diagram uses a convention where supported nodes are filled-in points, whereas unsupported nodes are hollow circles with associated displacement arrows. It is possible that a node can be stationary in one direction but not the other. For example, a roller would be free to move in the x direction but not the y. Professional FEM software handles a much wider variety of boundary conditions, and standard icons are used in the industry to depict these. The goal here is simplicity over sophistication.
The function trussGraphicsNodes does most of the work of
mapping the various types of nodes onto the specific graphics element.
The complexity of the code is managed by judicious use of patterns and
replacement rules. Some of the scaling and text placement was largely
determined by trial and error, so you may need to tweak these settings
for your own application or add additional code to help generalize the
solution.
As before, once the infrastructure is in place, the diagram is easy to create by simply asking the model for the "diagram" property.
Other important properties are axialStrain, axialStress, and axialForce. These will be implemented to
return all or specific values for a specified element.
There are many books and online resources that cover FEM. For example, the theory relevant to truss structures can be found at Jason Midkiff’s Virginia Tech science and engineering website: http://bit.ly/32BUq1 .
If you are looking for books with a Mathematica focus, look no further than Fundamental Finite Element Analysis and Applications: With Mathematica and Matlab Computations (John Wiley) and—if you are really into FEM— Advanced Topics in Finite Element Analysis of Structures: With Mathematica and MATLAB Computations (John Wiley), both by M. Asghar Bhutti. The code in these books is pre-version 6, but I found few incompatibilities.